

|
QBIPDFName:
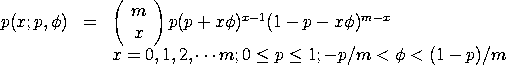
with p,
The quasi-binomial type I distribution is used to model
Bernoulli trials. The parameter p denotes the initial
probability of success, m denotes the number of Bernoulli
trials, and
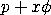
<SUBSET/EXCEPT/FOR qualification> where <x> is a positive integer variable, number, or parameter; <p> is a number, parameter, or variable in the range (0,1) that specifies the first shape parameter; <phi> is a number, parameter, or variable that specifies the second shape parameter; <m> is a number, parameter, or variable that specifies the third shape parameter; <y> is a variable or a parameter (depending on what <x> is) where the computed quasi binomial type I pdf value is stored; and where the <SUBSET/EXCEPT/FOR qualification> is optional.
LET Y = QBIPDF(X,0.7,0.01,20) PLOT QBIPDF(X,0.3,0.005,20) FOR X = 0 1 20
LET P = <value> LET PHI = <value> LET Y = QUASI BINOMIAL TYPE I RANDOM NUMBERS FOR I = 1 1 N
QUASI BINOMIAL TYPE I PROBABILITY PLOT Y
QUASI BINOMIAL TYPE I CHI-SQUARE GOODNESS OF FIT Y
In fitting the quasi-binomial type I distribution to data,
we typically assume that the number of trials, m, is
fixed and known and we then estimate p and
To obtain the maximum likelihood estimates of p and
QUASI BINOMIAL TYPE I MAXIMUM LIKELIHOOD Y QUASI BINOMIAL TYPE I MAXIMUM LIKELIHOOD Y2 X2 For unbinned data, the maximum likelihood estimates are the solutions to the equations:
![SUM[i=1 to N][(X(i)*(X(i) - 1)/(p + X(i)*PHI) -
SUM[i=1 to N][(M - X(i))/(1 - P - X(i)*PHI)] = 0](eqns/qbiml1.gif)
For binned data, the equations become
![SUM[i=1 to k][n(i)*(i-1)*i/(p+i*phi)] -
SUM[i=1 to k][n(i)*(m-1)*i/(1-p-i*phi)] = 0](eqns/qbiml3.gif)
with k, n, and nx denoting the number of classes, the total sample size, and the frequency of the xth class, respectively. These equations are known to have multiple solutions, so good starting values are required. By default, we use the starting values recommended by Consul and Famoye
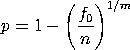
with f0 denoting the frequency of the
class x = 0 and
Alternatively, you can specify your own starting values by entering the commands
LET PHISTART = <value> Consul and Famoye give formulas for the Fisher information matrix (the inverse of the parameter variance-covariance matrix). They also give simplified formulas for the special cases m = 1, 2, or 3.
You can generate estimates of p and
LET P1 = <value> LET P2 = <value> LET PHI1 = <value> LET PHI2 = <value> QUASI BINOMIAL TYPE I KS PLOT Y QUASI BINOMIAL TYPE I KS PLOT Y2 X2 QUASI BINOMIAL TYPE I KS PLOT Y3 XLOW XHIGH QUASI BINOMIAL TYPE I PPCC PLOT Y QUASI BINOMIAL TYPE I PPCC PLOT Y2 X2 QUASI BINOMIAL TYPE I PPCC PLOT Y3 XLOW XHIGH The default values of p1 and p2 are 0.05 and 0.95, respectively. The default values for phi1 and phi2 are phi1 = -p1/m and phi2 = (1-p1)/m. Due to the discrete nature of the percent point function for discrete distributions, the ppcc plot will not be smooth. For that reason, if there is sufficient sample size the KS PLOT (i.e., the minimum chi-square value) is typically preferred. However, it may sometimes be useful to perform one iteration of the PPCC PLOT to obtain a rough idea of an appropriate neighborhood for the shape parameters since the minimum chi-square statistic can generate extremely large values for non-optimal values of the shape parameters. Also, since the data is integer values, one of the binned forms is preferred for these commands.
title size 3
tic label size 3
label size 3
legend size 3
height 3
x1label displacement 12
y1label displacement 15
.
multiplot corner coordinates 0 0 100 95
multiplot scale factor 2
label case asis
title case asis
case asis
tic offset units screen
tic offset 3 3
title displacement 2
y1label Probability Mass
x1label X
.
ylimits 0 1
major ytic mark number 6
minor ytic mark number 3
xlimits 0 20
line blank
spike on
.
multiplot 2 2
.
title P = 0.3, Phi = 0.01, M = 20
plot qbipdf(x,0.3,0.01,20) for x = 1 1 20
.
title P = 0.3, Phi = -0.01, M = 20
let phi = -0.01
plot qbipdf(x,0.3,phi,20) for x = 1 1 20
.
title P = 0.7, Phi = 0.01, M = 20
plot qbipdf(x,0.7,0.01,20) for x = 1 1 20
.
title P = 0.7, Phi = -0.01, M = 20
let phi = -0.01
plot qbipdf(x,0.7,phi,20) for x = 1 1 20
.
end of multiplot
.
justification center
move 50 97
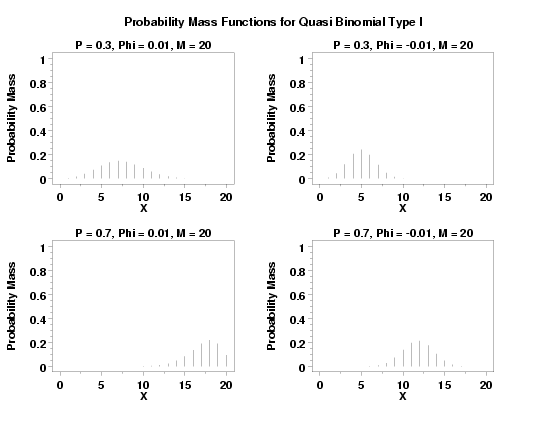
text Probability Mass Functions for Quasi Binomial Type I
Program 2:
let p = 0.7
let phi = 0.01
let m = 20
let psave = p
let phisave = phi
let y = quasi binomial type I rand numb for i = 1 1 500
.
let y3 xlow xhigh = integer frequency table y
class lower -0.5
class width 1
class upper 20.5
let y2 x2 = binned y
.
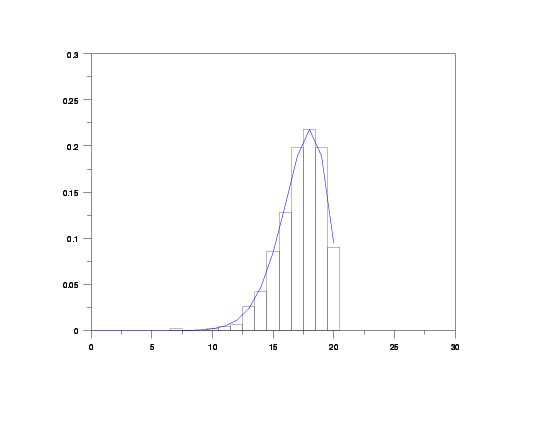
relative hist y2 x2
multiplot 2 2 4
limits freeze
pre-erase off
line color blue
plot qbipdf(x,p,phi,m) for x = 0 1 20
limits
pre-erase on
line color black
.
quasi binomial type I mle y
let p = pml
let phi = phiml
.
quasi binomial type I chi-square goodness of fit y3 xlow xhigh
.
char x
line bl
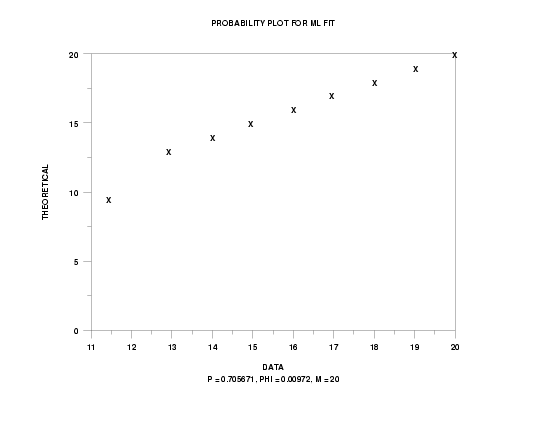
quasi binomial type I probability plot y3 xlow xhigh
.
char bl
line so
let p1 = 0.5
let p2 = 0.9
let a1 = (1 - p1)/m
let a2 = (1 - p2)/m
let phi1 = 0
let phi2 = max(a1,a2)
quasi binomial type I ks plot y3 xlow xhigh
let p = shape1
let phi = shape2
quasi binomial type I chi-square goodness of fit y3 xlow xhigh
QUASI BINOMIAL TYPE I PARAMETER ESTIMATION:
NUMBER OF OBSERVATIONS = 500
SAMPLE MEAN = 17.26600
SAMPLE STANDARD DEVIATION = 1.901177
SAMPLE MINIMUM = 7.000000
SAMPLE MAXIMUM = 20.00000
ZERO-CLASS FREQUENCY: = 0.000000
USER-SPECIFIED VALUE FOR M = 20.00000
MAXIMUM LIKELIHOOD:
ESTIMATE OF P = 0.7056712
ESTIMATE OF PHI = 0.9720844E-02
THE COMPUTED VALUE OF THE CONSTANT P = 0.7056712E+00
THE COMPUTED VALUE OF THE CONSTANT PHI = 0.9720844E-02
CHI-SQUARED GOODNESS-OF-FIT TEST
NULL HYPOTHESIS H0: DISTRIBUTION FITS THE DATA
ALTERNATE HYPOTHESIS HA: DISTRIBUTION DOES NOT FIT THE DATA
DISTRIBUTION: QUASI BINOMIAL TYPE I
SAMPLE:
NUMBER OF OBSERVATIONS = 500
NUMBER OF NON-EMPTY CELLS = 9
NUMBER OF PARAMETERS USED = 3
TEST:
CHI-SQUARED TEST STATISTIC = 1.331433
DEGREES OF FREEDOM = 5
CHI-SQUARED CDF VALUE = 0.068336
ALPHA LEVEL CUTOFF CONCLUSION
10% 9.23636 ACCEPT H0
5% 11.07050 ACCEPT H0
1% 15.08628 ACCEPT H0
Date created: 8/23/2006 |